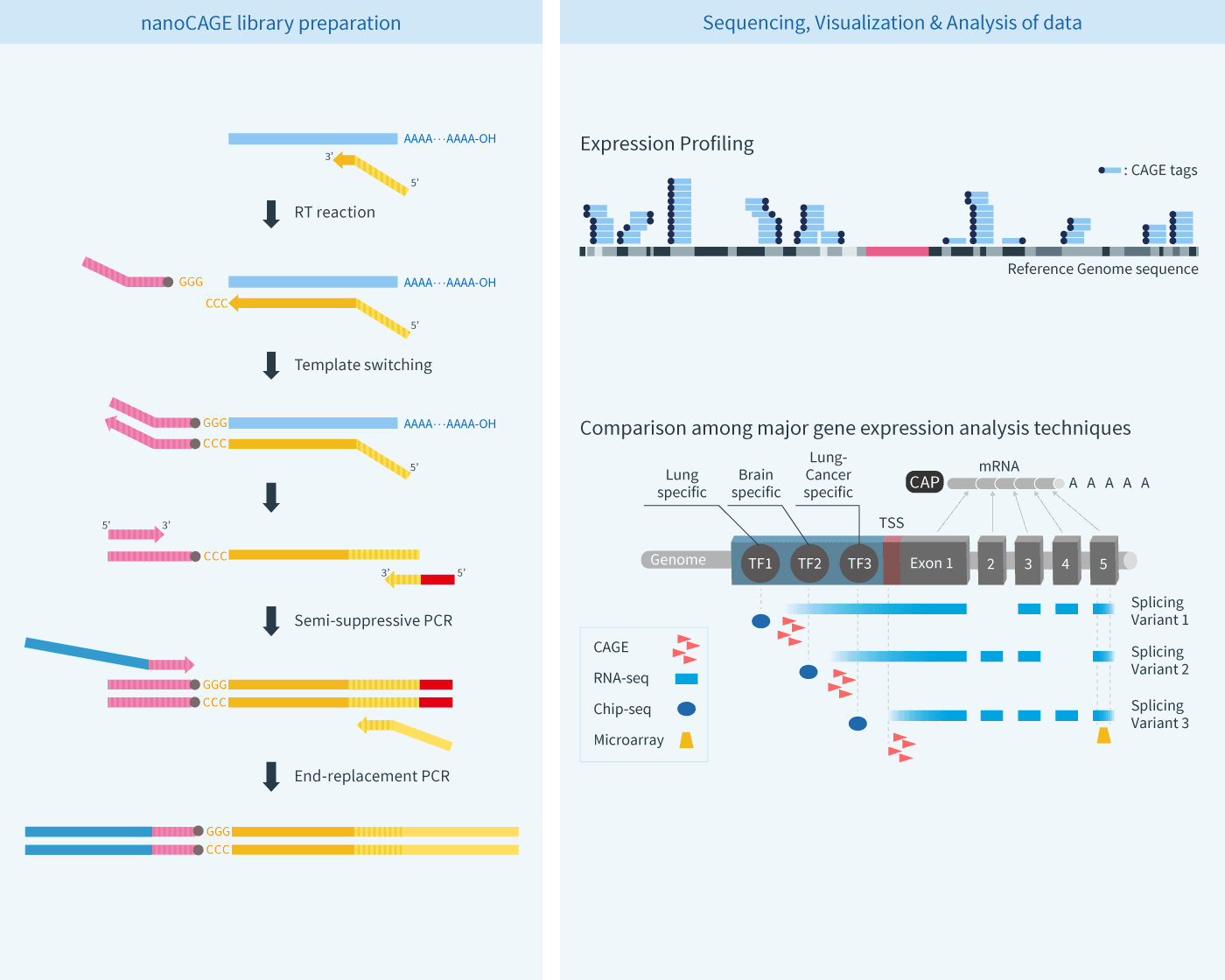
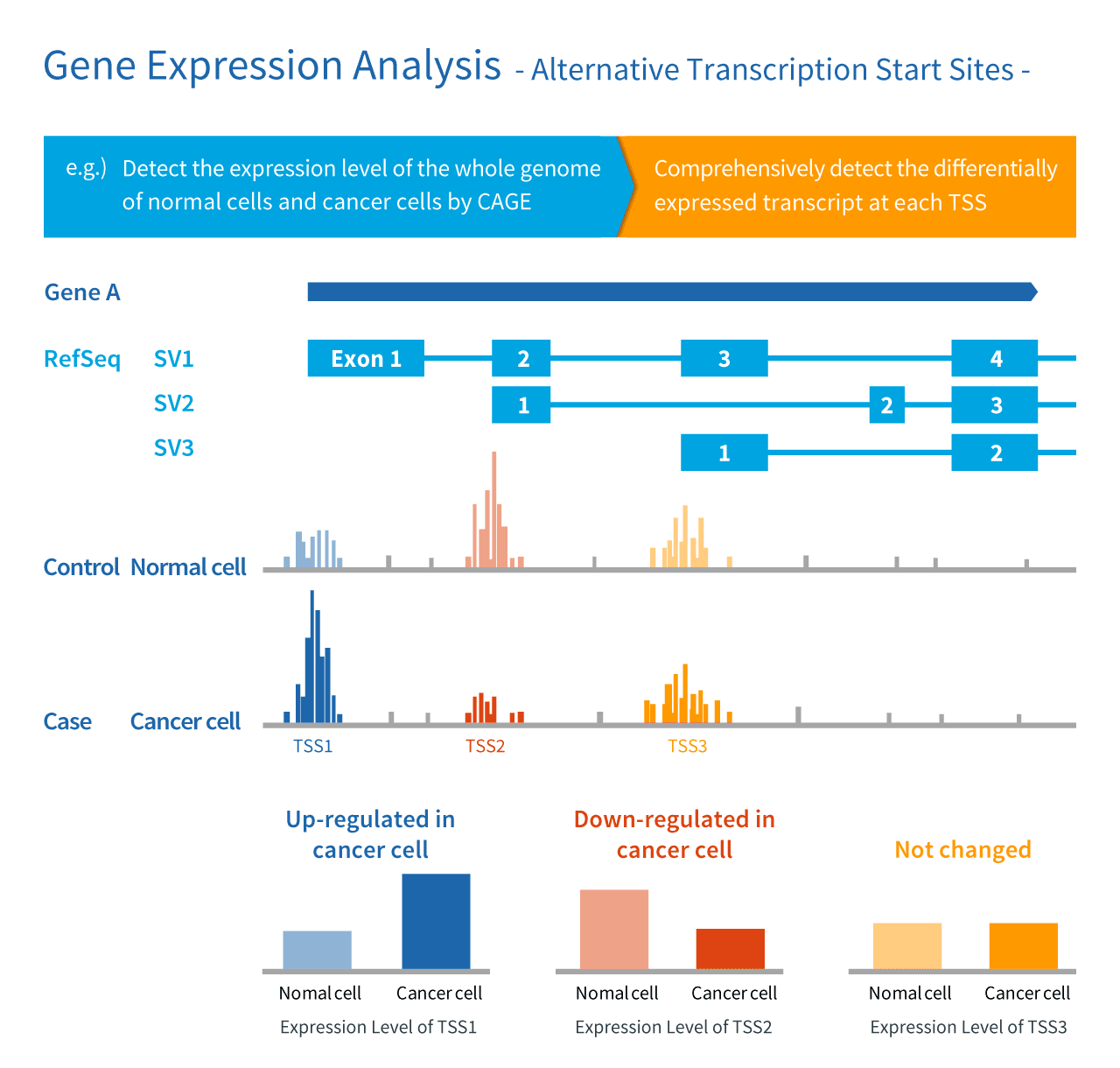
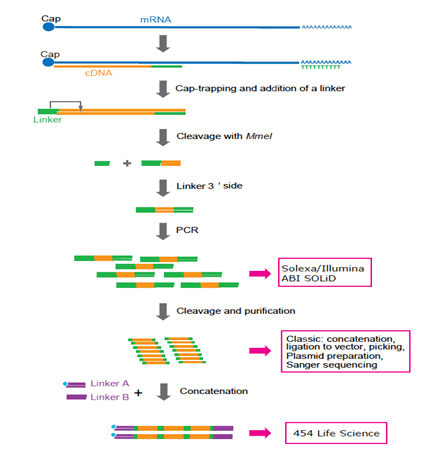
Frontiers | Extensive Variation in Gene Expression is Revealed in 13 Fertility-Related Genes Using RNA-Seq, ISO-Seq, and CAGE-Seq From Brahman Cattle

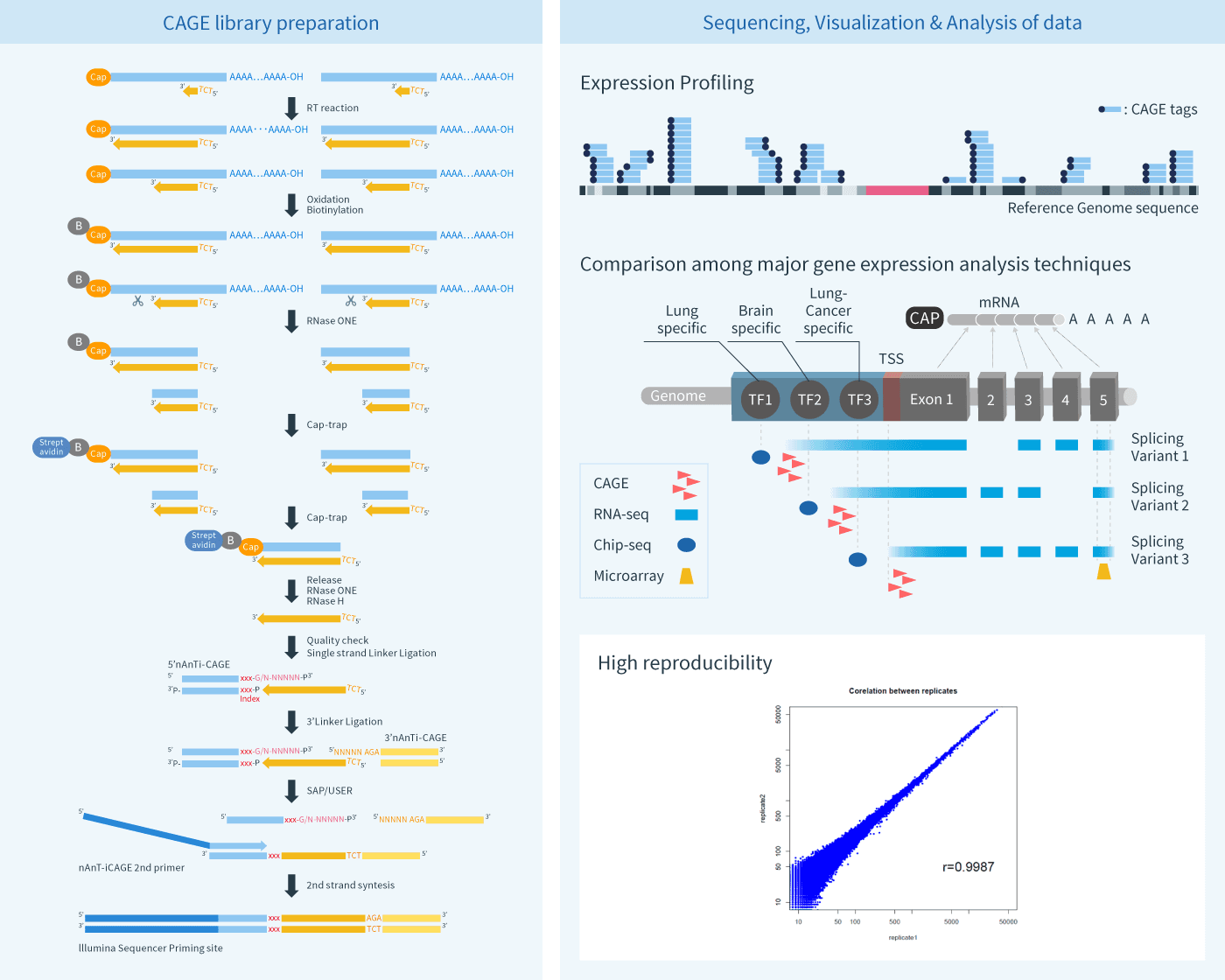
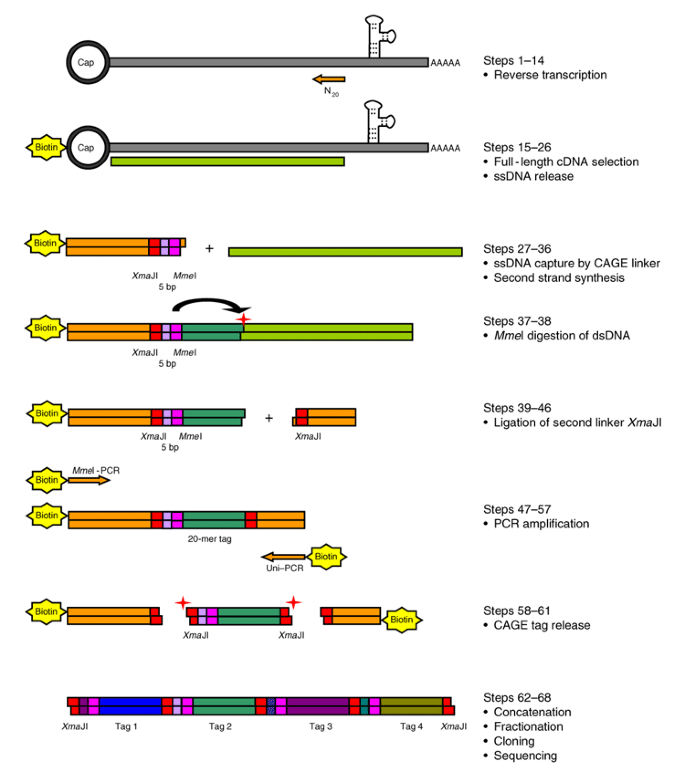
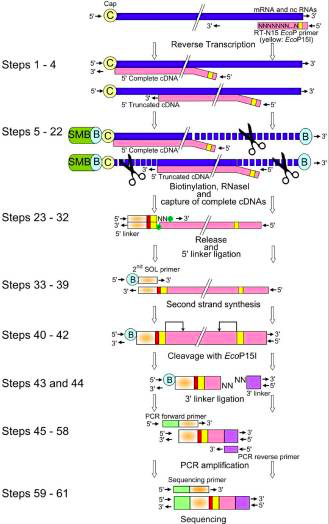
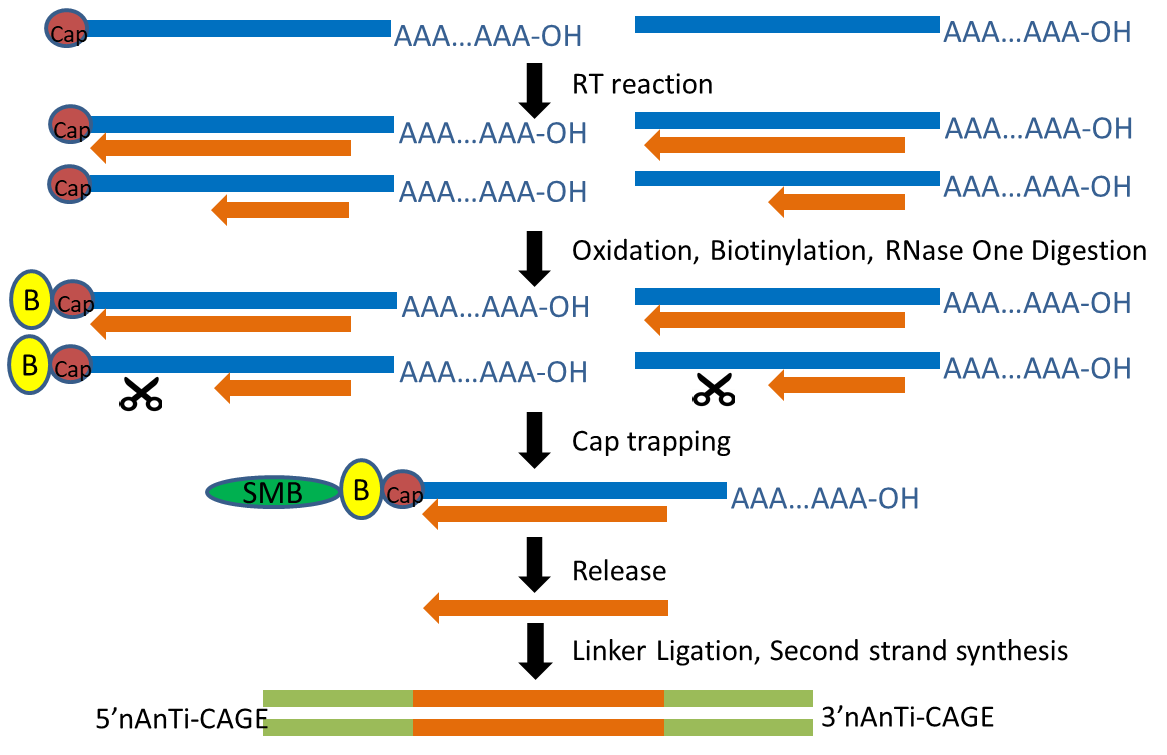
CAGE-seq characterization of alternative promoter usage and full-length... | Download Scientific Diagram

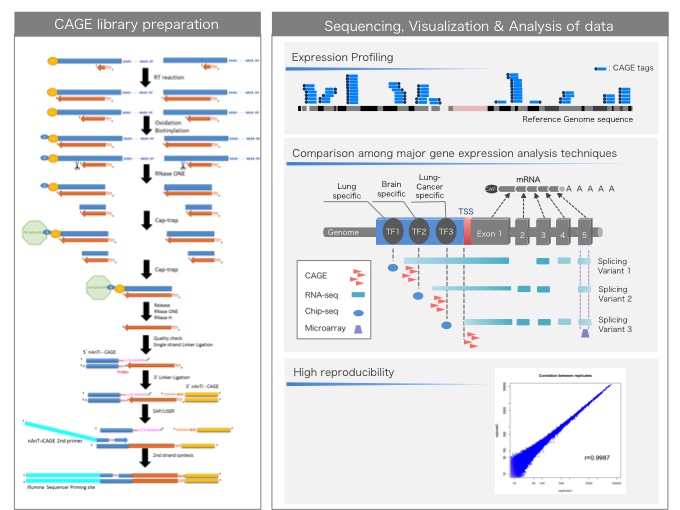
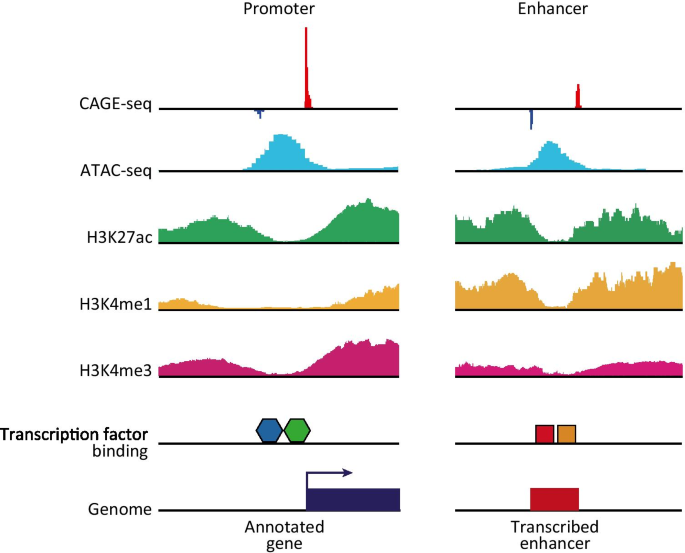
NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

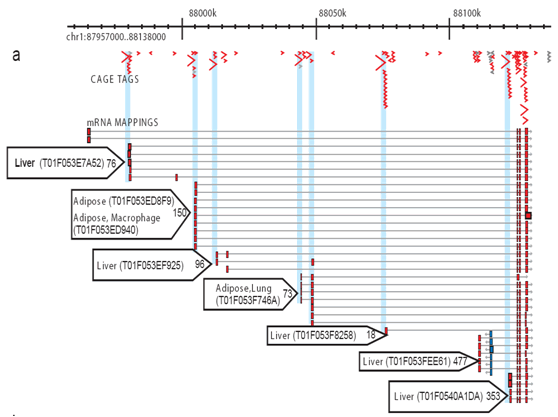
Schematic diagram of CAGE and RNA-Seq read coverage for a gene with two... | Download Scientific Diagram
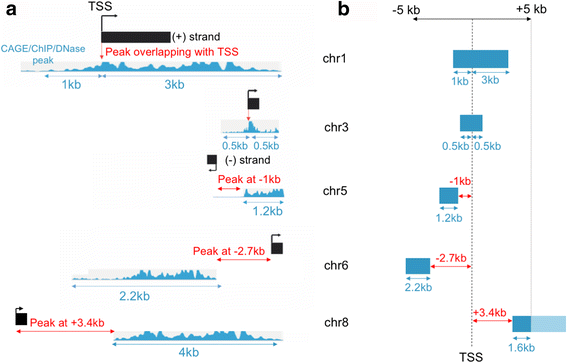
Zipper plot: visualizing transcriptional activity of genomic regions | BMC Bioinformatics | Full Text
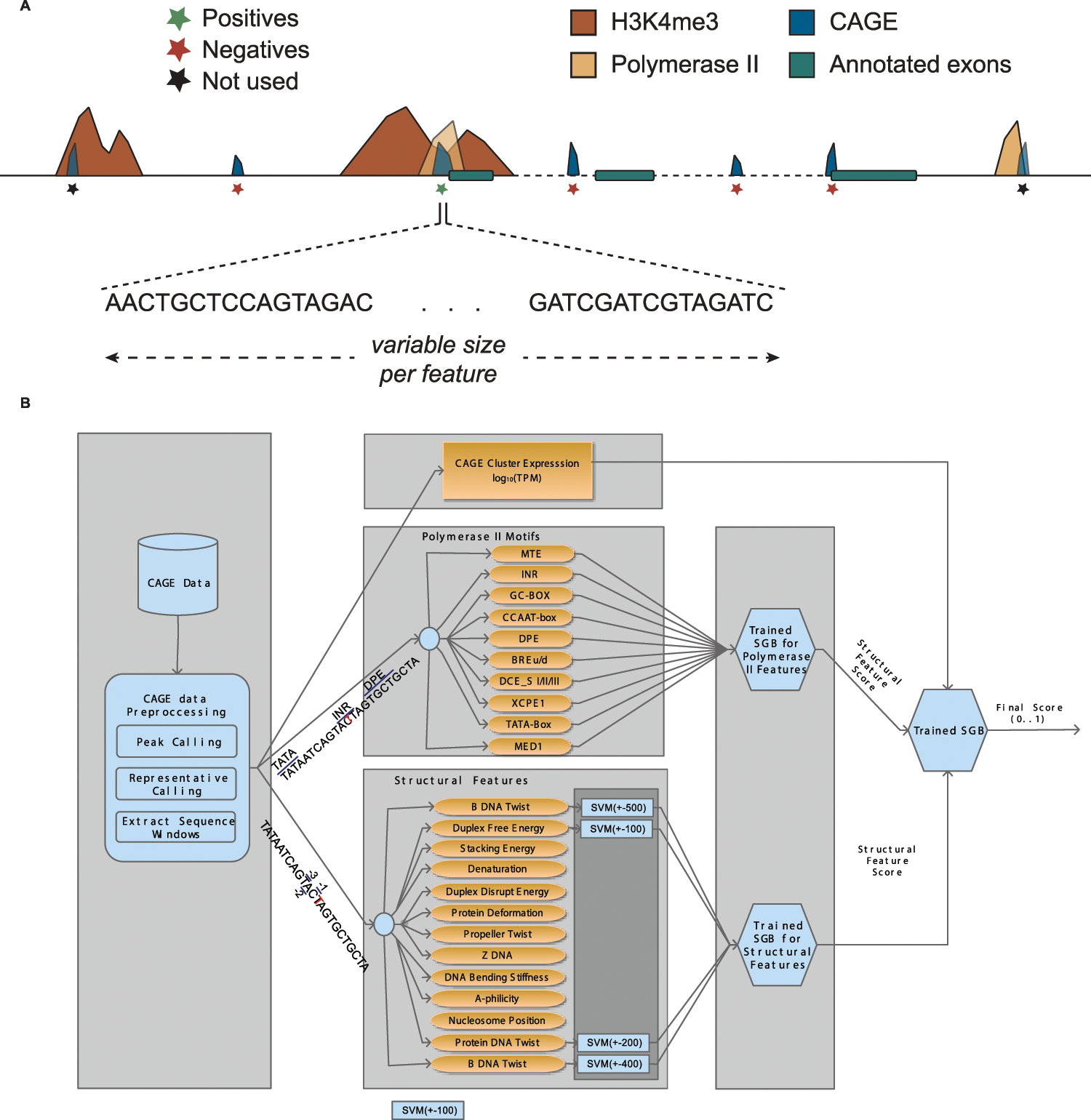
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports